Task 6
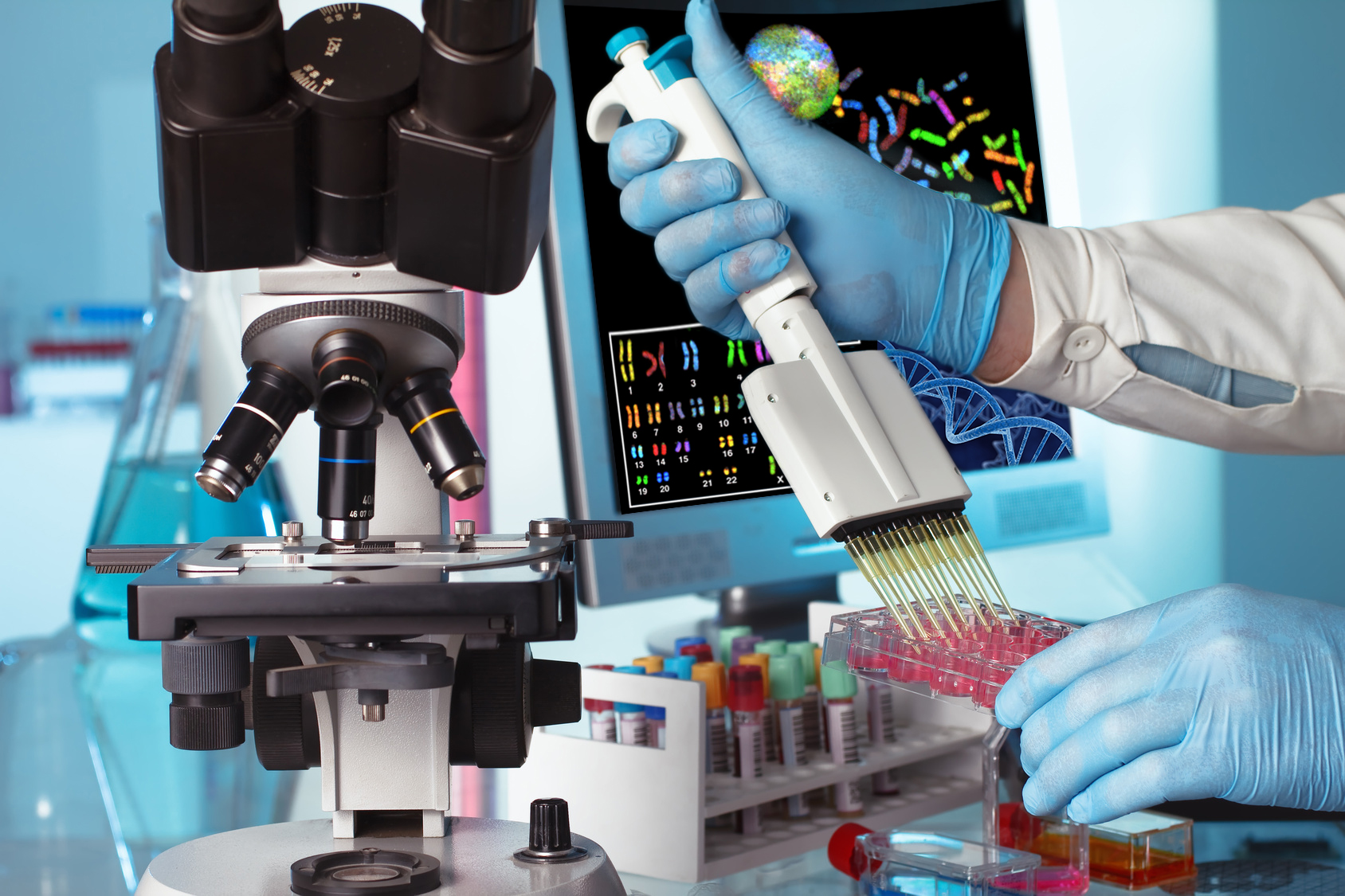

This task aims to identify genetic markers that are directly associated with phenotypic variation of resistance to parasites. Identification of genetic markers among the 600.000 SNPs of the Ovine HD BeadChip array will be based on described approaches using open-source whole genome association analysis toolsets. Parasite resistance traits will be analyzed using Genome-wide Efficient Mixed Model Analysis for Association Studies (GEMMA). To correct for population stratification, an analysis of variance will be performed, where gender, lambing season, birth rank (single, multiple) and age of dam will be fitted as fixed effects and age as a linear co-variable.
Post GWAS analysis will focus the search for genes close to the significant SNPs. Since most studies on sheep resistance to nematodes utilized microsatellite markers, the respective base pair positions will be searched for comparison with the positions found with Illumina Ovine HD Beadchip. In addition, the Bos taurus genome will be searched for homologies and genes at flanking regions of the SNPs of interest. With this information, biological pathways will be searched for each gene.
Institutions involved and members of the team in this task:
INIAV
Expected results:
- Identification of significant SNP markers correlated with phenotypic resistance traits
- Identification of candidate genes
- Identification of biological pathways involved in resistance
- Identification of tissue types expressing candidate genes